Tutorial, Part 4¶
In this part of the tutorial, we look at the queries that are defined
in project.xml and the use of “rewrites” to aggregate the
results in different ways.
4.0 Queries¶
The queries identified in the project file (or in an external file) determine which results are extracted from the GCAM database for each run of the model, and thus determine which subsequent steps (computing differences, creating charts) can be performed.
GCAM uses an XML-based database for which queries are likewise composed in XML. The database is managed by the java-based ModelInterface program provided in the GCAM distribution. There is also a standard file called “Main_Queries.xml” that is used by ModelInterface to provide interactive access to these queries.
Pygcam executes queries by creating XML query files and invoking the ModelInterface
program in “batch” (non-interactive) mode to generate CSV files. You can craft query
files by hand, or you can use pre-existing ones in Main_Queries.xml or some other
file with custom queries.
The queries themselves can be extracted on-the-fly from these files by specifying
the location of the XML file in the configuration variable GCAM.QueryPath and
referencing the desired query by its defined “title”. (See the
query sub-command and the pygcam.query API documentation
for more information.) In general, there is little need to create individual query
files; anything you can run in ModelInterface can be run by pygcam as well.
Queries can be run several ways in GCAM:
- If an XML database is written to disk (the default), queries can be run on the database using the ModelInterface.jar file, which is used by the query sub-command.
- If the XML database is written to disk, GCAM can run the queries before it exits, using the same mechanism as in the option above.
- Since v4.3, GCAM can write its XML database to memory only, in which case it must be queried from within GCAM since the database will no longer exist after GCAM exits. This is particularly useful in large ensemble (e.g., Monte Carlo simulation) runs where you want to extract some data but don’t need to keep the large databases around.
Two configuration file parameters control this behavior. The variables and
their default values are shown below. Add these to your .pygcam.cfg file
with appropriate True or False values to configure GCAM as you wish.
# Setting ``GCAM.InMemoryDatabase`` to ``True`` forces ``GCAM.RunQueriesInGCAM``
# to be ``True`` since there is no other way to run queries in this case.
GCAM.InMemoryDatabase = False
GCAM.RunQueriesInGCAM = False
Note
Using the in-memory database substantially increases GCAM’s memory footprint, particularly since version 5.0, so it may be impractical to use this feature in some cases.
4.1 Processing of query definitions¶
When the project.xml file is read, the <queries> element is saved to
a temporary file, the pathname of which is stored in the variable given by the
varName attribute. In the case above, the pathname is stored in queryXmlFile.
The stored filename can be accessed in command steps using curly braces, i.e.,
{queryXmlFile}. The query and and diff sub-commands both understand
the format of this file. The query sub-command obviously runs the queries as
indicated, whereas the diff command uses the query names to identify the
resulting CSV files that should be compared. Examples of the <step> elements
using the temporary query file are as follows:
<step name="query" runFor="policy">@query -o {batchDir} -w {scenarioDir} -s {scenario} -Q "{queryPath}" -q "{queryXmlFile}"</step>
<step name="diff" runFor="policy">@diff -D {sandboxDir} -y {years} -Y {shockYear} -q "{queryXmlFile}" -i {baseline} {scenario}</step>
Note that the double-quotes around {queryXmlFile} are necessary only if the pathname
contains blanks; using them is good “defensive programming” practice.
4.2 Rewrite sets¶
Standard GCAM XML queries can define “rewrites” which modify the values of chosen data elements to allow them to be aggregated. For example, you can aggregate all values of CornAEZ01, CornAEZ02, …, CornAEZ18 to be returned simply as “Corn”.
In pygcam this idea is taken a step further by allowing you to define reusable,
named “rewrite sets” that can be applied to queries named in the project file.
For example, if you are working with a particular
regional aggregation, you can define this aggregation once in a rewrites.xml file
and reference the name of the rewrite set when specifying queries in project.xml.
See rewrite sets for more information.
Defining queries with rewrites¶
The following example rewriteSets.xml file is copied into new projects:
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 46 47 48 49 50 51 52 53 54 55 56 57 58 59 60 61 62 63 64 65 66 67 68 69 70 71 72 73 74 75 76 77 78 79 80 81 82 83 84 85 86 87 88 89 90 91 92 93 94 95 96 97 98 99 100 101 102 103 104 105 106 107 108 109 110 111 112 | <?xml version="1.0"?> <rewriteSets xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance" xsi:noNamespaceSchemaLocation="rewriteSets-schema.xsd"> <rewriteSet name="liquidFuels" level="technology" append-values="false"> <rewrite from="cellulosic ethanol" to="Cellulosic ethanol"/> <rewrite from="corn ethanol" to="Corn ethanol"/> <rewrite from="sugar cane ethanol" to="Sugar cane ethanol"/> <rewrite from="cellulosic ethanol CCS level 1" to="Cellulosic ethanol"/> <rewrite from="cellulosic ethanol CCS level 2" to="Cellulosic ethanol"/> <rewrite from="FT biofuels" to="FT biofuels"/> <rewrite from="biodiesel" to="Oilcrop biodiesel"/> <rewrite from="FT biofuels CCS level 1" to="FT biofuels"/> <rewrite from="FT biofuels CCS level 2" to="FT biofuels"/> <rewrite from="coal to liquids" to="CTL"/> <rewrite from="coal to liquids CCS level 1" to="CTL"/> <rewrite from="coal to liquids CCS level 2" to="CTL"/> <rewrite from="crude oil refining" to="Oil refining"/> <rewrite from="oil refining" to="Oil refining"/> <rewrite from="gas to liquids" to="GTL"/> <rewrite from="unconventional oil refining" to="Oil refining"/> </rewriteSet> <rewriteSet name="eightRegions" level="region" append-values="true"> <rewrite from="USA" to="United States"/> <rewrite from="Brazil" to="Brazil"/> <rewrite from="Canada" to="Rest of World"/> <rewrite from="China" to="China"/> <rewrite from="Africa_Eastern" to="Africa"/> <rewrite from="Africa_Northern" to="Africa"/> <rewrite from="Africa_Southern" to="Africa"/> <rewrite from="Africa_Western" to="Africa"/> <rewrite from="Japan" to="Rest of Asia"/> <rewrite from="South Korea" to="Rest of Asia"/> <rewrite from="India" to="Rest of Asia"/> <rewrite from="Central America and Caribbean" to="Rest of South America"/> <rewrite from="Central Asia" to="Rest of Asia"/> <rewrite from="EU-12" to="Europe Union 27"/> <rewrite from="EU-15" to="Europe Union 27"/> <rewrite from="Europe_Eastern" to="Rest of World"/> <rewrite from="Europe_Non_EU" to="Rest of World"/> <rewrite from="European Free Trade Association" to="Rest of World"/> <rewrite from="Indonesia" to="Rest of Asia"/> <rewrite from="Mexico" to="Rest of South America"/> <rewrite from="Middle East" to="Rest of World"/> <rewrite from="Pakistan" to="Rest of Asia"/> <rewrite from="Russia" to="Rest of World"/> <rewrite from="South Africa" to="Africa"/> <rewrite from="South America_Northern" to="Rest of South America"/> <rewrite from="South America_Southern" to="Rest of South America"/> <rewrite from="South Asia" to="Rest of Asia"/> <rewrite from="Southeast Asia" to="Rest of Asia"/> <rewrite from="Taiwan" to="Rest of Asia"/> <rewrite from="Argentina" to="Rest of South America"/> <rewrite from="Colombia" to="Rest of South America"/> <rewrite from="Australia_NZ" to="Rest of Asia"/> </rewriteSet> <rewriteSet name="food" level="input"> <rewrite from="Corn" to="Grains"/> <rewrite from="FiberCrop" to="Other"/> <rewrite from="MiscCrop" to="Other"/> <rewrite from="OilCrop" to="Other"/> <rewrite from="OtherGrain" to="Grains"/> <rewrite from="PalmFruit" to="Other"/> <rewrite from="Rice" to="Grains"/> <rewrite from="Root_Tuber" to="Other"/> <rewrite from="SugarCrop" to="Other"/> <rewrite from="Wheat" to="Grains"/> <rewrite from="regional beef" to="Meat"/> <rewrite from="Dairy" to="Meat"/> <rewrite from="OtherMeat_Fish" to="Meat"/> <rewrite from="Pork" to="Meat"/> <rewrite from="Poultry" to="Meat"/> <rewrite from="SheepGoat" to="Meat"/> </rewriteSet> <rewriteSet name="landCover" level="LandLeaf" byAEZ="true"> <rewrite from="biomass" to="Biomass"/> <rewrite from="Corn" to="Cropland"/> <rewrite from="eucalyptus" to="Cropland"/> <rewrite from="FiberCrop" to="Cropland"/> <rewrite from="FodderGrass" to="Cropland"/> <rewrite from="FodderHerb" to="Cropland"/> <rewrite from="Forest" to="Forest (managed)"/> <rewrite from="Grassland" to="Grass"/> <rewrite from="Jatropha" to="Cropland"/> <rewrite from="miscanthus" to="Biomass"/> <rewrite from="MiscCrop" to="Cropland"/> <rewrite from="OilCrop" to="Cropland"/> <rewrite from="OtherArableLand" to="Cropland"/> <rewrite from="OtherGrain" to="Cropland"/> <rewrite from="PalmFruit" to="Cropland"/> <rewrite from="Pasture" to="Pasture (grazed)"/> <rewrite from="ProtectedGrassland" to="Other arable land"/> <rewrite from="ProtectedShrubland" to="Other arable land"/> <rewrite from="ProtectedUnmanagedForest" to="Forest (unmanaged)"/> <rewrite from="ProtectedUnmanagedPasture" to="Pasture (other)"/> <rewrite from="Rice" to="Cropland"/> <rewrite from="RockIceDesert" to="Other land"/> <rewrite from="Root_Tuber" to="Cropland"/> <rewrite from="Shrubland" to="Other arable land"/> <rewrite from="SugarCrop" to="Cropland"/> <rewrite from="Tundra" to="Other land"/> <rewrite from="UnmanagedForest" to="Forest (unmanaged)"/> <rewrite from="UnmanagedPasture" to="Pasture (other)"/> <rewrite from="UrbanLand" to="Other land"/> <rewrite from="Wheat" to="Cropland"/> <rewrite from="willow" to="Cropland"/> <rewrite from="SugarcaneEthanol" to="Cropland"/> </rewriteSet> </rewriteSets> |
We can reference any of these sets in the <queries> section of the project.xml
file. We can define a list of rewrite sets to apply by default to all queries, and
we can define rewrites to apply to individual queries (as well as opt out of the
default rewrites in any individual query.)
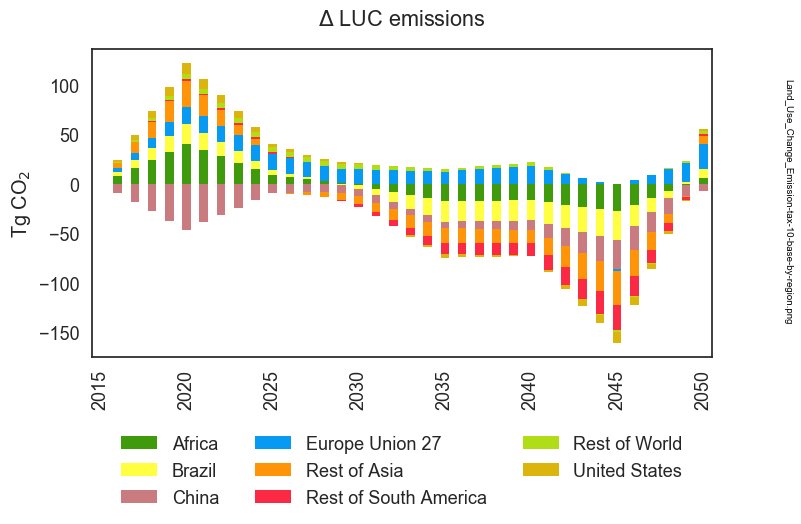
Let’s now use the pre-defined “eightRegions” set to aggregate the 32 regions to
simplify the plot of Land Use Change Emissions we’ve been working on. To do this,
we change the line for this query in project.xml from
<query name="Land_Use_Change_Emission"/>
to:
<query name="Land_Use_Change_Emission">
<rewriter name="eightRegions"/>
</query>
We then need to rerun the queries for both the baseline and policy scenarios, recompute the differences, and re-generate the plots. We can do that with this command:
$ gt run -s query,diff,plotDiff -S base,tax-10
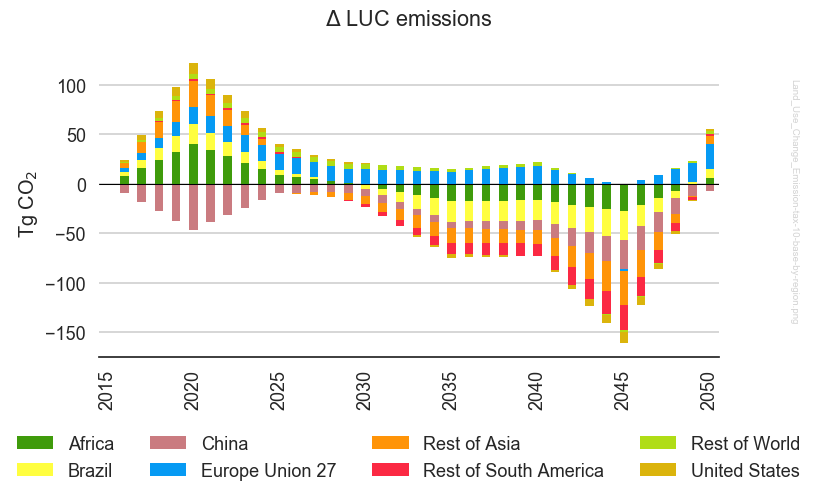
This results in the following figure:
or, if we restore the original aesthetic choices, we have this: